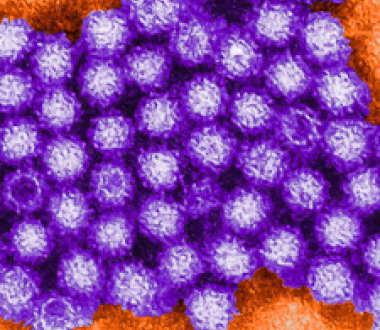
Norovirus is a primary cause of acute gastroenteritis and foodborne disease in the United States where, according to the CDC, it is responsible for "19 - 21 million cases of acute gastroenteritis, 1.7 - 1.9 million outpatient visits and 400,000 emergency room visits per year. It also contributes to between 56,000 - 71,000 hospitalizations and 570 - 800 deaths per year, mostly among young children and the elderly."
The name CaliciNet is a reference to the virus family Caliciviridae which includes the genus Norovirus. Noroviruses are currently classified into six genogroups. Nearly all human norovirus infections are caused by strains belonging to genogroups I and II. Noroviruses are further subdivided into genotypes. For example, pandemic GII.4 strains belong to genogroup II, genotype 4.
CaliciNet is a network of federal, state and local public health laboratories established in 2009 to track norovirus outbreaks, trace causative strains to the source, standardize the typing of strains, and identify new strains. These are important tasks because the emergence of a new strain can result in a 50% increase in the number of cases.
In the United States there are only five CaliciNet Outbreak Support Centers. As of 2014, 33 laboratories were forwarding their specimens to these Support Centers for testing. Participation as a CaliciNet Outbreak Support Center requires certification in a range of laboratory methods and bioinformatics skills in addition to bi-annual participation in proficiency testing provided by the CDC. As a CaliciNet Outbreak Support Center, Dr. Patrick Bryant’s Enteric Virology Laboratory at the Wadsworth Center is performing norovirus typing for several states including Rhode Island, West Virginia, Connecticut, Maine and Pennsylvania. Testing and reporting are complex processes which consist of the following steps:
As a CaliciNet Outbreak Support Center, Dr. Patrick Bryant’s Enteric Virology Laboratory at the Wadsworth Center is performing norovirus typing for several states including Rhode Island, West Virginia, Connecticut, Maine and Pennsylvania. Testing and reporting are complex processes which consist of the following steps:
1. Perform conventional RT-PCR to amplify specific regions.
2. Sequence these regions. (This step is performed by the Wadsworth Center Applied Genomic Technologies Core).
3. Analyze the sequences to determine the specific norovirus strain responsible.
4. Transmit sequence and associated epidemiological data through a secure pipeline to the National CaliciNet Database.
Next, uploaded data is compared to existing sequence data in the database, occasionally identifying a new strain. For example, during the 2015-2016 season, Wadsworth Center’s Enteric Virology Laboratory identified a new strain of norovirus from Asia. The GII.17 Kawasaki virus was identified in samples from several suspected foodborne outbreaks in Pennsylvania and New York.
Advanced scientific methods of identifying strains, evaluating relatedness and tracking them to the source facilitate the implementation of appropriate control measures which minimize outbreak spread and reduce the risk of additional cases.
This publication was supported by Cooperative Agreement # 56400-200-801-17-01 funded by the Centers for Disease Control and Prevention through the Association of Public Health Laboratories. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the Centers for Disease Control and Prevention, the Department of Health and Human Services or the Association of Public Health Laboratories.