Services
Targeted (Amplicon) DNA Sequencing
Targeted DNA sequencing allows the researcher to utilize the specificity of PCR in order to target the genes of their choosing, including 16S metagenomic studies. Targeted DNA sequencing provides the ability to achieve deeper sequencing coverage in order to identify those genes expressed at lower levels that may possibly have been missed by other sequencing methods.
Microbial Whole-Genome Sequencing (WGS)
Microbial sequencing is the whole-genome sequencing of a bacterial chromosome. It can assist in the discovery of genetic variations that support the designing of antimicrobial compounds, vaccines, and even engineered microbes for industrial applications.
Bacterial and viral typing through WGS is used in the accurate and fast identification and discrimination of strains. Bacterial and viral typing can also assist in outbreak identification, surveillance, resistance-typing and the understanding of transmission, pathogenesis, and evolutionary relationships of the target. Often specific isolates can be sequenced rapidly using next-generation sequencing techniques.
Instruments
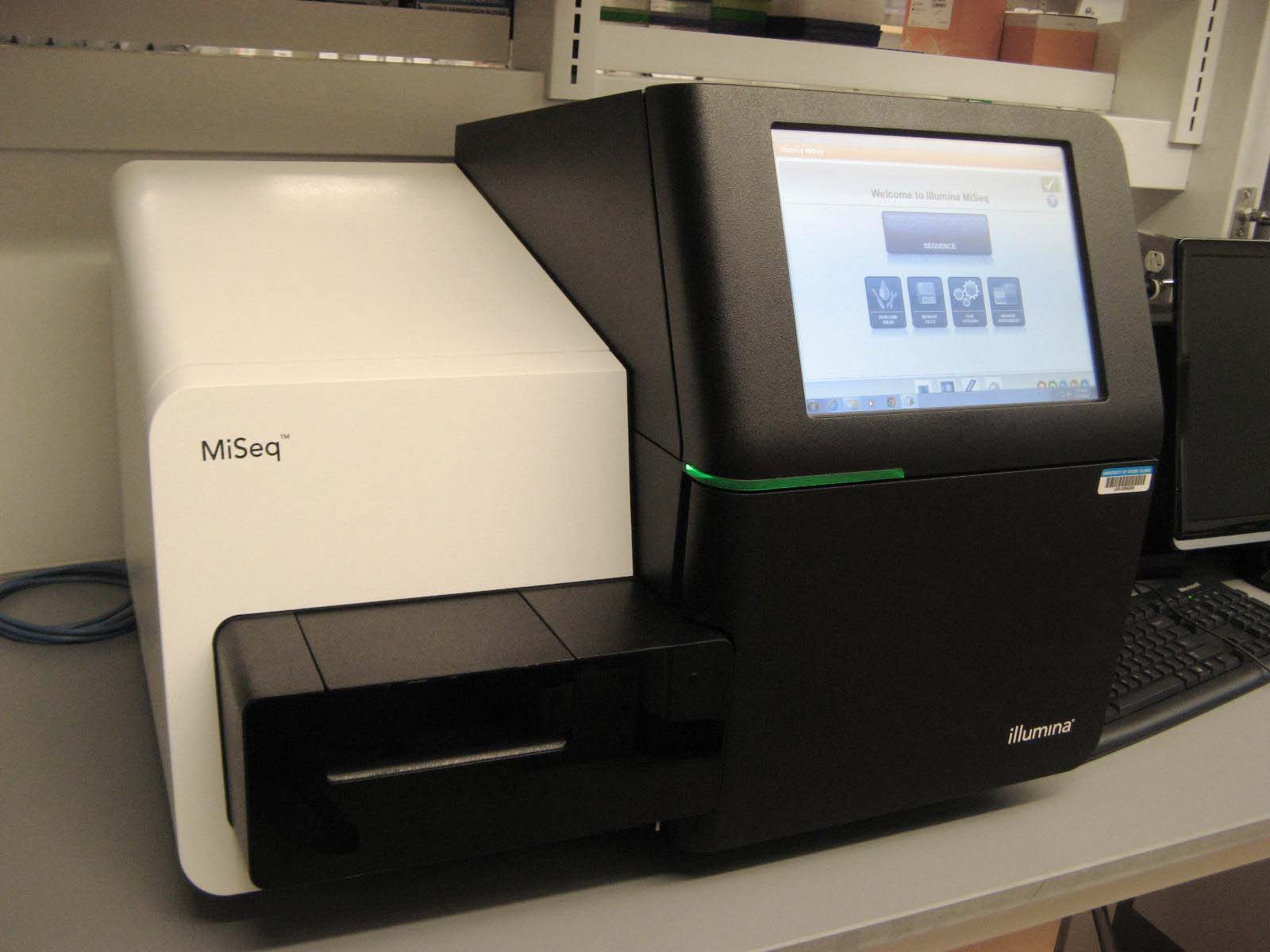

Illumina MiSeq
The MiSeq is a benchtop sequencer optimized for speed (as little as 6 hours for 50bp single reads) and long reads (2x300 bp). It is fully compatible with libraries made for the Illumina HiSeq. Because of its fast turn-around time, this instrument is often used for testing library protocols before sequencing on the HiSeq. But because of its unique read length and throughput, it is also a good instrument for data production for small genomes or amplicon sequencing projects. We support and make libraries for DNA-Seq, RNA-Seq, 16S metagenomics, and Tru-Seq custom amplicon panels. We also support other amplicon sequencing.
MiSeq Specifications
Cluster Generation and Sequencing - MiSeq Reagent Kit v2
| Read Length | Approximate Total Time* | Output | Quality Scores†† |
|---|---|---|---|
| 1 x 36 bp | 4 hours | 540-610 Mb | >90% bases higher than Q30 |
| 2 x 25 bp | 5.5 hours | 750-850 Mb | >90% bases higher than Q30 |
| 2 x 150 bp | 24 hours | 4.5-5.1 Gb | >80% bases higher than Q30 |
| 2 x 250 bp | 39 hours | 7.5-8.5 Gb | >75% bases higher than Q30 |
MiSeq Reagent Kit v2 Reads Passing Filter†: single reads = 12-15 M; paired-end reads = 24-30 M
Cluster Generation and Sequencing - MiSeq Reagent Kit v3
| Read Length | Approximate Total Time* | Output | Quality Scores†† |
|---|---|---|---|
| 2 x 75 bp | 20 hours | 3.3 - 3.8 Gb | >85% bases higher than Q30 |
| 2 x 300 bp | 55 hours | 13.2 - 15 Gb | >70% bases higher than Q30 |
MiSeq Reagent Kit v3 Reads Passing Filter†: single reads = 22-25 M; paired-end reads = 44-50 M
*Total time includes cluster generation, sequencing, and basecalling on a MiSeq System enabled with dual-surface scanning.
†Install specifications based on Illumina PhiX control library at supported cluster densities (between 467-583 k/mm2 clusters passing filter for v2 chemistry and 727-827 k/mm2 clusters passing filter for v3 chemistry). Actual performance parameters may vary based on sample type, sample quality, and clusters passing filter.
††A quality score (Q-score) is a prediction of the probability of an error in base calling. The percentage of bases > Q30 is averaged across the entire run.
bp = base pairs, Mb = megabases, Gb = gigabases, M = millions.
Ion Torrent PGM
We offer sequencing services on the Ion Torrent PGM. This sequencer uses semiconductor chip technology. They simply sequence DNA by monitoring the hydrogen ions released every time a nucleotide is incorporated. There is no enzymatic cascade, no fluorescence, no chemiluminescence, etc. Three chips with different throughput capacity, shown below, are available: 314, 316, and 318 chips.
Ion Torrent PGM System Performance Specifications
| Ion 314 Chip v2 | Ion 316 Chip v2 | Ion 318 Chip v2 | ||
|---|---|---|---|---|
| Output | 200 base | 30-50 Mb | 300-600 Mb | 600 Mb-1 Gb |
| 400 base | 60-100 Mb | 600 Mb-1 Gb | 1.2-2 Gb | |
| Reads | 400-550 thousand | 2-3 million | 4-5.5 million | |
| Run Time | 200 base | 2.3 hours | 3.0 hours | 4.4 hours |
| 400 base | 3.7 hours | 4.9 hours | 7.3 hours |
Library Quality Control
To confirm the size range of the library, the AGTC uses an Agilent Bioanalyzer or Tapestation. The final concentration of each library is determined using the Qubit, prior to sequencing.

BioAnalyzer 2100
Qubit 2.0 Fluorometer

Tapestation 2200
Service Requests and Submission Instructions
To request next-generation sequencing services or for additional information, please email or call 518-486-4099 the Applied Genomic Technologies Core.